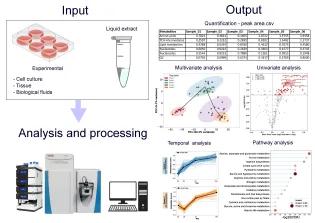
Metabolomics
Map which metabolites are present, how they change, and where carbon flows.
Semi-Targeted Profiling
Quantitative profiling using our in-house validated library of 425 metabolites, covering energy metabolism (glycolysis, TCA cycle), nucleotide metabolism (purines and pyrimidines), amino acid metabolism, and more. You get confident identification, not just masses.
Untargeted Profiling
High-resolution MS/MS acquisition searched against spectral libraries for broad, hypothesis-free discovery. Suitable when you want to find what changes without pre-defining the targets.
Isotope Tracing / Metabolic Flux Analysis
Track where carbon (¹³C), nitrogen (¹⁵N), or deuterium flows through metabolic networks. Understand which pathways are actively used, not just which metabolites are present. Ideal for studying metabolic reprogramming in cancer, immunity, or pharmacological intervention.